Biomedisa is a free, open-source, and user-friendly application designed for the segmentation of large 3D image datasets such as CT and MRI scans. It has been developed at The Australian National University CTLab. Biomedisa's smart interpolation of sparsely pre-segmented slices enables accurate semi-automated segmentation by considering the full underlying image data. In addition, Biomedisa supports deep learning workflows for fully automated segmentation across similar samples and structures. Biomedisa can be used both online and locally. It is compatible with widely used segmentation tools such as Amira/Avizo, and ImageJ/Fiji, and it also provides a 3D Slicer extension.
About Biomedisa
References and Links
Updated on 7 jan 2026 by Philipp
[Online App] Use Biomedisa online without installation. [Installation] Follow the installation instructions on GitHub. [Biomedisa] Nat. Commun. 11, 5577 (2020). [Deep Learning] PLoS Comput. Biol. 19, e1011529 (2023). [Smart Interpolation] Proc. SPIE 9784, 97842L (2016). [HEDI Hernia Repair] Commun. Med. 6, 68 (2026). [Particle Separation] Preprint at https://doi.org/10.48550/arXiv.2508.16224 (2025). [Antscan] Nat. Methods 23, 663–672 (2026). [Antscan Project] https://www.antscan.info [Biomedisa Database] https://biomedisa.info/gallery/
Antscan: A synchrotron micro-CT-based resource of ant diversity
Posted on 13 mar 2026 by Philipp
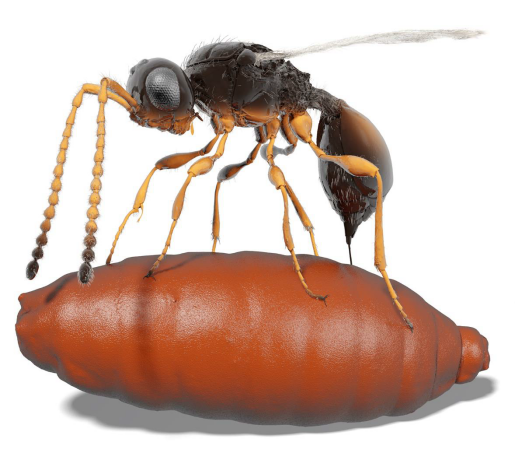
About the cover image: 3D ant models derived from synchrotron micro-CT and made available via Biomedisa. Ant species (clockwise from top): Paraponera clavata, Camponotus brutus, Daceton armigerum, Cephalotes clypeatus, Eciton hamatum, Discothyrea sexarticulata. Read the full publication. Image credit: Thomas van de Kamp, Karlsruhe Institute of Technology (KIT). Cover design: Thomas Phillips.
Large-scale library of 3D ant images
Posted on 6 mar 2026 by Philipp
Thousands of 3D ant scans generated using synchrotron X-ray microtomography are now available through the Biomedisa platform. Biomedisa hosts the interactive database and provides integrated AI tools for automated segmentation of the specimens. Read the full publication publication.
Scientists reconstruct the face of Little Foot, a 4 million-year-old human ancestor
Posted on 15 mar 2026 by Philipp
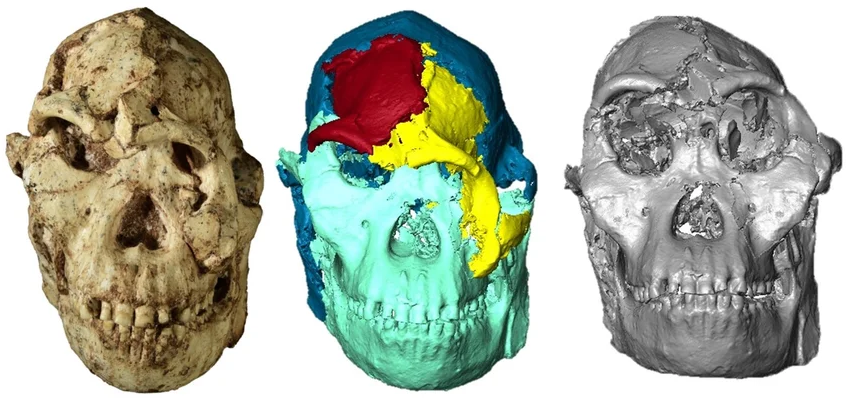
“Lucy” may be the most famous Australopithecus, but “Little Foot” is the most complete fossil ever discovered from the genus. The image shows a facial reconstruction of Little Foot (right), created from the original skull (left) and a digital replica (center) using Biomedisa. Read the full publication. Image credit: Amélie Beaudet, CNRS, Université de Poitiers. Sources: CNN Science and Scientific American.
Registrations are open for VIS2026
Posted on 3 feb 2026 by Philipp
VIS2026 marks Volume Imaging Australia’s third symposium! We invite you to join us to highlight 3D volumetric imaging and analysis across all branches of science. The program continues to focus on four main themes—LIGHT, ELECTRON, X-RAY, and CORRELATIVE volume imaging—including innovations in visualisation and data analysis shared across these modalities. Register here.
Command-line Usage
Posted on 12 Jan 2026 by Philipp
Perform interpolation-based segmentation using sparse labels. Additional options are described in the Smart Interpolation documentation.
python -m biomedisa.interpolation <image_file> <label_file>
Arguments:
<image_file>– Path to the input image (e.g..tif,.nrrd, or.am)<label_file>– Path to the corresponding label file (e.g..tif,.nrrd, or.am)--allaxis– Enables interpolation along all spatial axes
Train a deep learning model using annotated image data. Additional options are described in the Deep Learning documentation.
python -m biomedisa.deeplearning <train_images> <train_labels> -t
Arguments:
<train_images>– Directory containing training images (e.g..tif,.nrrd, or.am)<train_labels>– Directory containing training labels (e.g..tif,.nrrd, or.am)--val_images– Directory containing validation images--val_labels– Directory containing validation labels
Apply a trained model to generate segmentations for new images. Additional options are described in the Deep Learning documentation.
python -m biomedisa.deeplearning <image_file> <model_file>
Arguments:
<image_file>– Path to the input image (e.g..tif,.nrrd, or.am)<model_file>– Path to a trained model (e.g..h5,.pth, including pretrained Biomedisa or Segment Anything Models for object separation)--mask– Path to a binary mask defining the region of interest for separation
Python Examples
Posted on 22 aug 2023 by Philipp
Perform interpolation-based segmentation using sparse labels. Additional options are described in the Smart Interpolation documentation.
from biomedisa.features.biomedisa_helper import load_data, save_data
from biomedisa.interpolation import smart_interpolation
# load data
img, _ = load_data('trigonopterus.tif')
labels, header = load_data('labels.trigonopterus_smart.am')
# smart interpolation with optional smoothing result
results = smart_interpolation(img, labels, smooth=100)
# get results
regular_result = results['regular']
smoothed = results['smooth']
# save results
save_data('final.trigonopterus.am', regular_result, header=header)
save_data('final.trigonopterus.smooth.am', smoothed, header=header)
Train a deep learning model using annotated image data. Additional options are described in the Deep Learning documentation.
from biomedisa.features.biomedisa_helper import load_data
from biomedisa.deeplearning import deep_learning
# load image data
img1, _ = load_data('Head1.am')
img2, _ = load_data('Head2.am')
img_data = [img1, img2]
# load label data
label1, _ = load_data('Head1.labels.am')
label2, header, ext = load_data('Head2.labels.am',
return_extension=True)
label_data = [label1, label2]
# load validation data (optional)
img3, _ = load_data('Head3.am')
img4, _ = load_data('Head4.am')
label3, _ = load_data('Head3.labels.am')
label4, _ = load_data('Head4.labels.am')
val_img_data = [img3, img4]
val_label_data = [label3, label4]
# deep learning
deep_learning(img_data, label_data, train=True, batch_size=12,
val_img_data=val_img_data, val_label_data=val_label_data,
header=header, extension=ext, path_to_model='honeybees.h5')
Apply a trained model to generate segmentations for new images. Additional options are described in the Deep Learning documentation.
from biomedisa.features.biomedisa_helper import load_data, save_data
from biomedisa.deeplearning import deep_learning
# load data
img, _ = load_data('Head5.am')
# deep learning
results = deep_learning(img, predict=True,
path_to_model='honeybees.h5', batch_size=6)
# save result
save_data('final.Head5.am', results['regular'], results['header'])
Biomedisa Segmentation Featured on the Cover of the Journal of Insect Science
Posted on 4 mar 2026 by Philipp
A Biomedisa-generated segmentation result has been selected as the cover image of the Journal of Insect Science. The image depicts the male head of Nasonia giraulti, showing the genomandibular gland and mandibular musculature. Read the full publication. Image credit: István Mikó, University of New Hampshire.
HEDI: Biomechanical Evaluation and Visualisation in Incisional Hernia Repair
Updated on 7 jan 2026 by Philipp
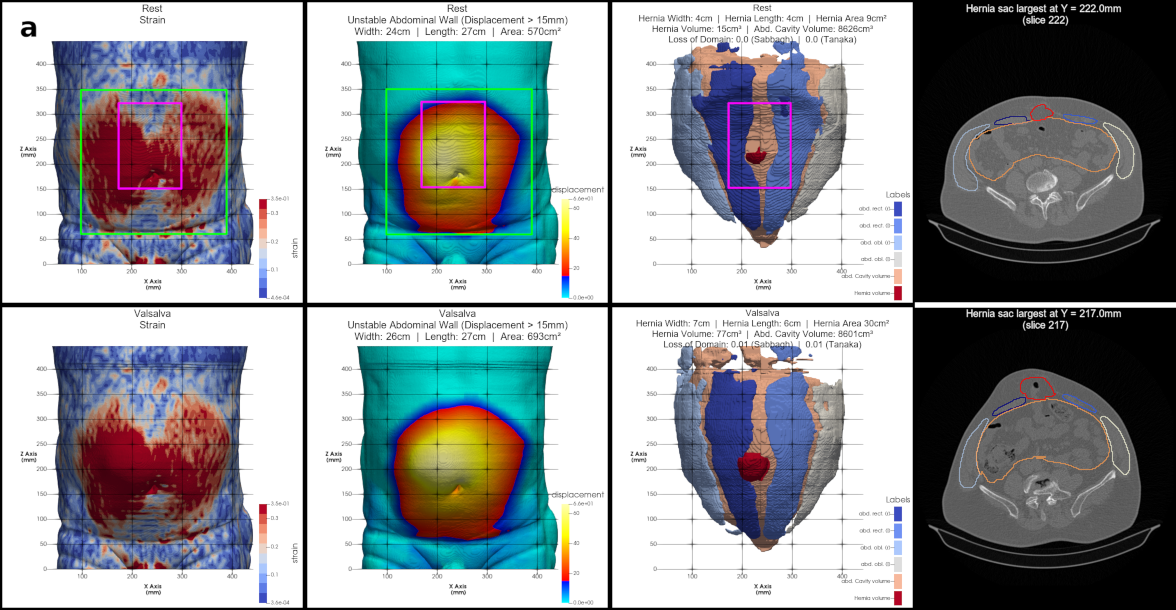
HEDI is a tool that uses computed tomography with Valsalva maneuver to detect and assess hernia size, volume, and abdominal wall instability. Our first clinical application of HEDI in the preoperative evaluation of 31 patients shows significantly improved success rates compared to reported rates, with all patients remaining pain-free and experiencing no hernia recurrence after three years of follow-up. Check out the paper published in Communications Medicine.
Smart Interpolation: Multi-Platform Support
Posted on 13 aug 2025 by Philipp
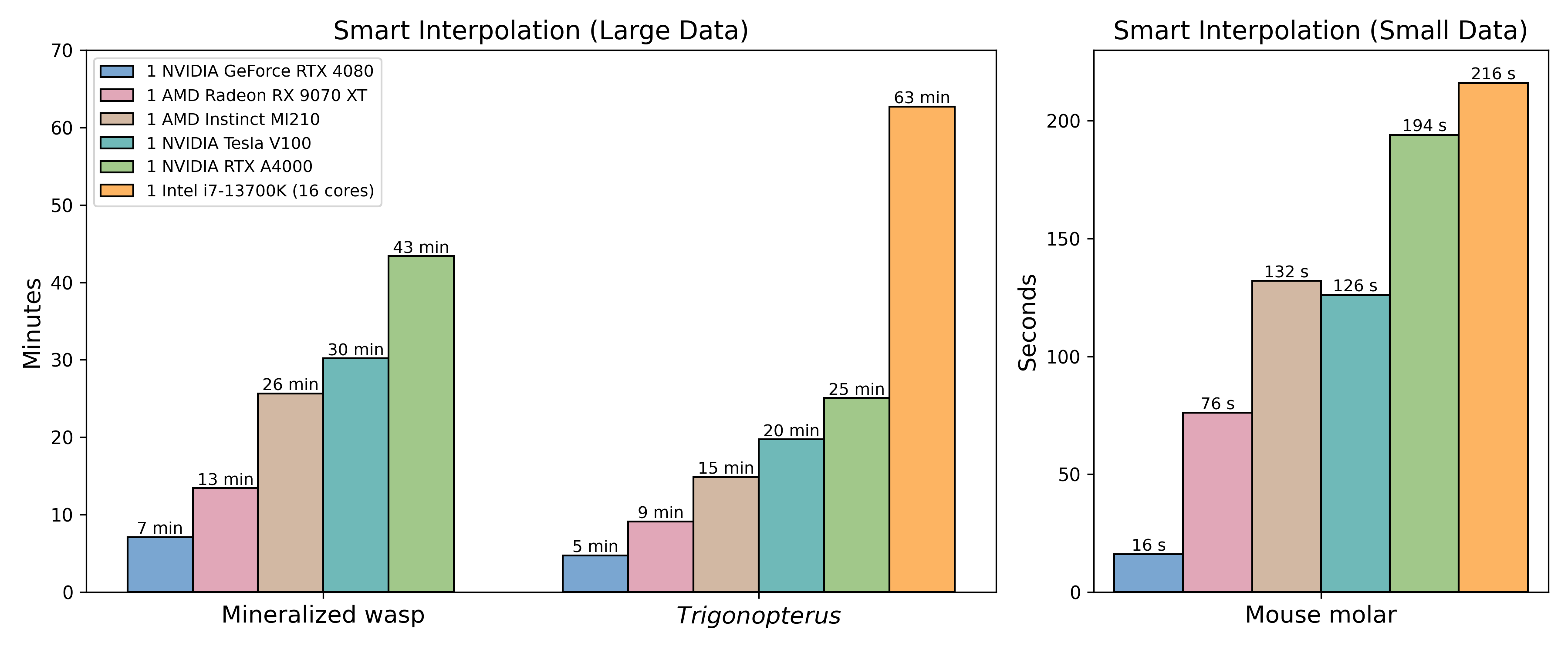
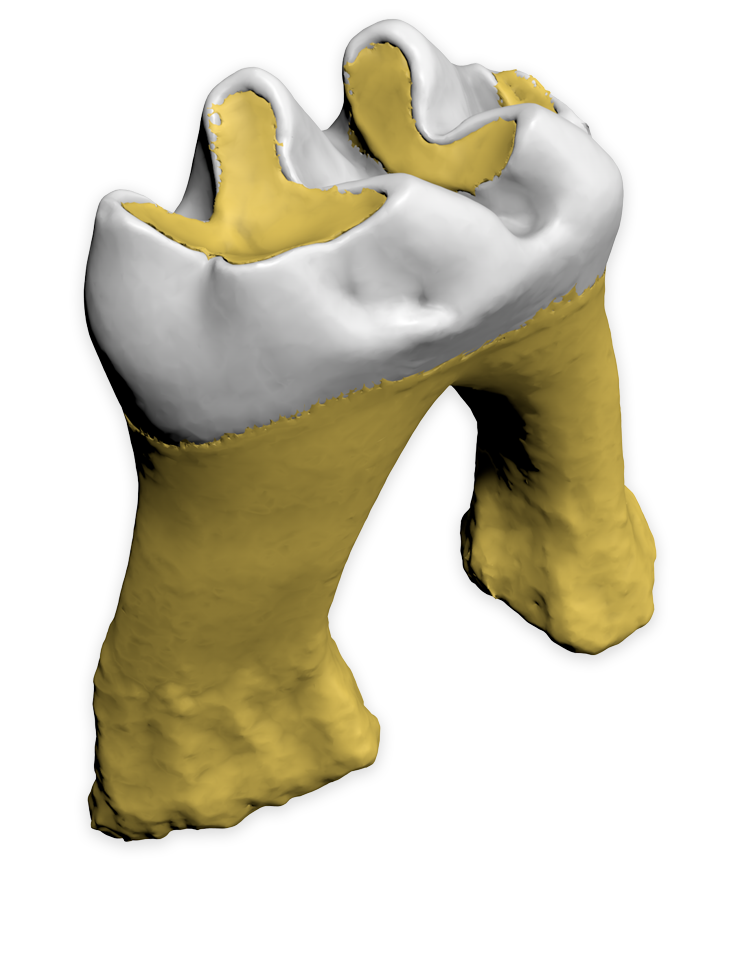
Biomedisa's Smart Interpolation works across NVIDIA, AMD, and Intel GPUs, as well as Intel CPUs. To get started, follow the installation instructions. By default, Biomedisa automatically chooses the best available platform for your system. You can also manually specify a platform, for example with ‑p=cuda, ‑p=opencl_NVIDIA_GPU, or ‑p=opencl_Intel_CPU. Tests displayed were run on datasets from the gallery. Mineralized wasp 1077×992×2553 voxels, every 20th slice pre-segmented, Trigonopterus 1497×734×1117 voxels, and Mouse molar 413×553×413 voxels.
Tutorial: How to use Smart Interpolation in 3D Slicer
Posted on 7 mar 2025 by Philipp
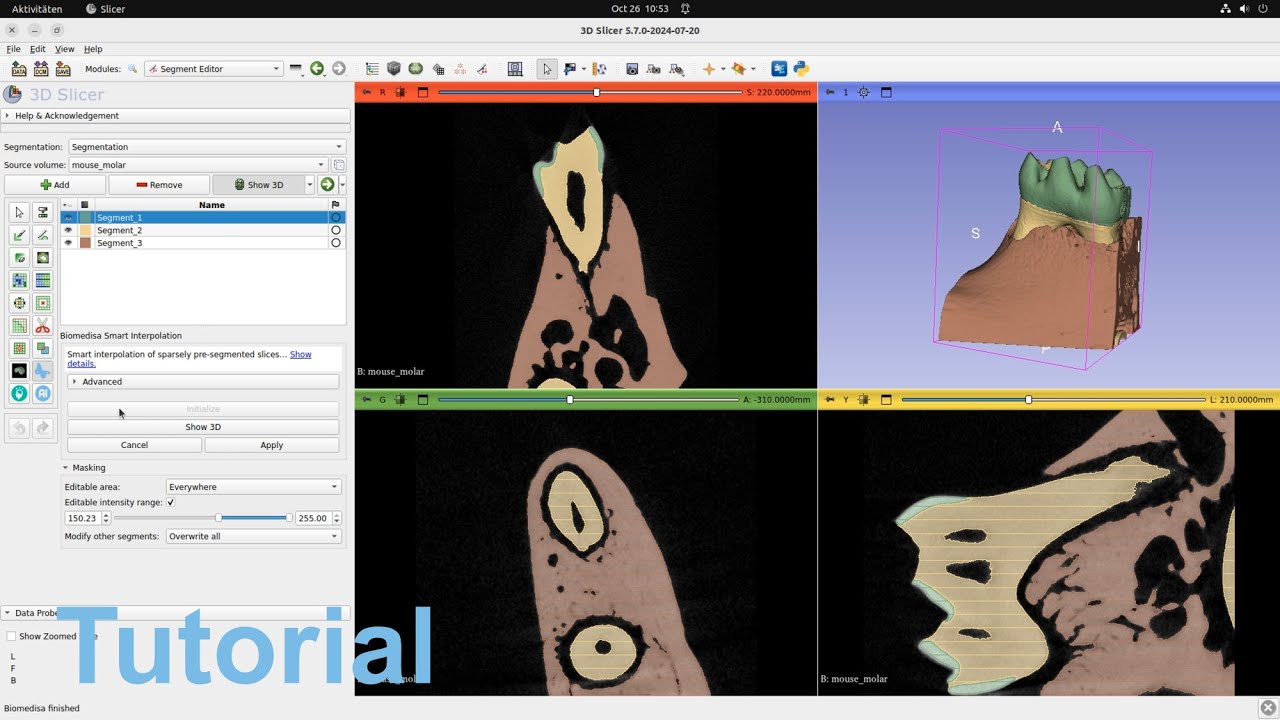
In this short video, we demonstrate how to use Biomedisa’s Smart Interpolation in 3D Slicer. Follow the installation instructions on GitHub to get started.
Webinar: Using Biomedisa and its 3D Slicer extension for biological image segmentation
Posted on 11 sep 2024 by Philipp
In this webinar presented by Volume Imaging Australia and Microscopy Australia, you learn how to use Biomedisa in combination with 3D Slicer, both freely available open-source platforms for biological and medical image analysis. We begin by using Biomedisa’s smart interpolation to semi-automatically create training data. This data will then be used to train Biomedisa’s deep neural network for automated segmentation, demonstrated through example cases such as mouse molar teeth from micro-CT scans and mitochondria in electron microscopy images.
Large-scale analysis using Biomedisa's Deep Learning
Posted on 3 oct 2023 by Philipp
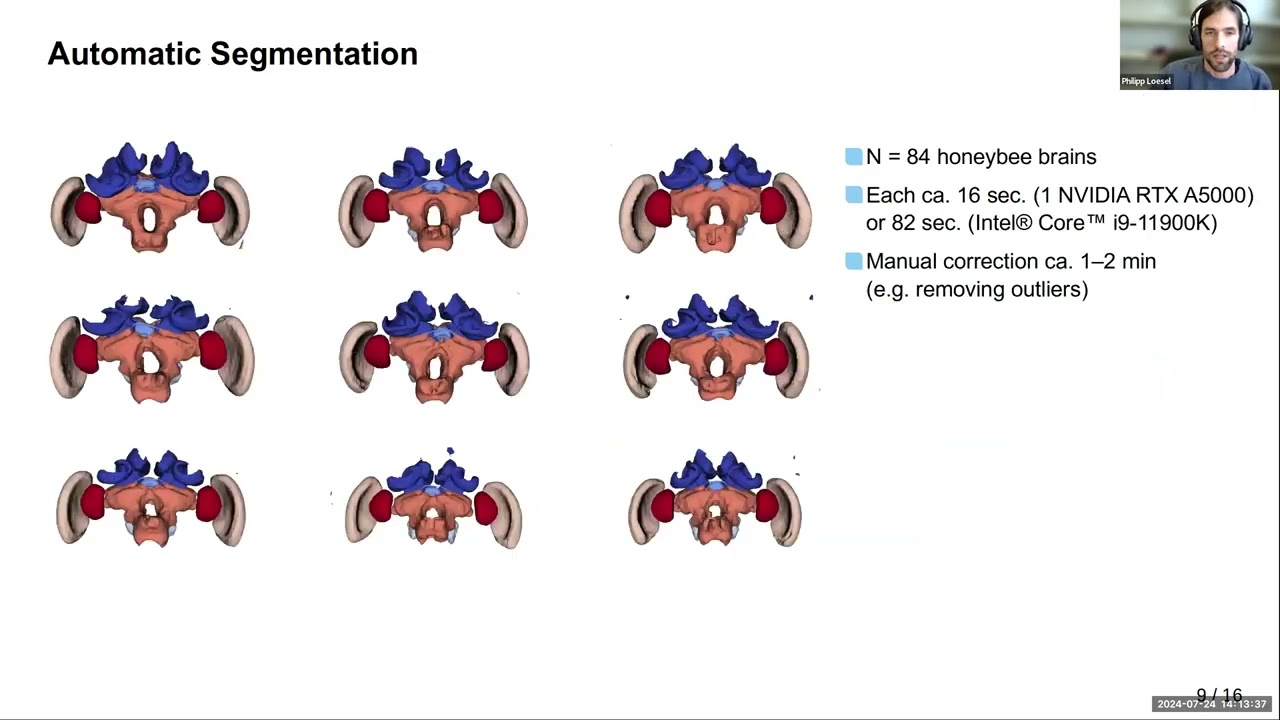
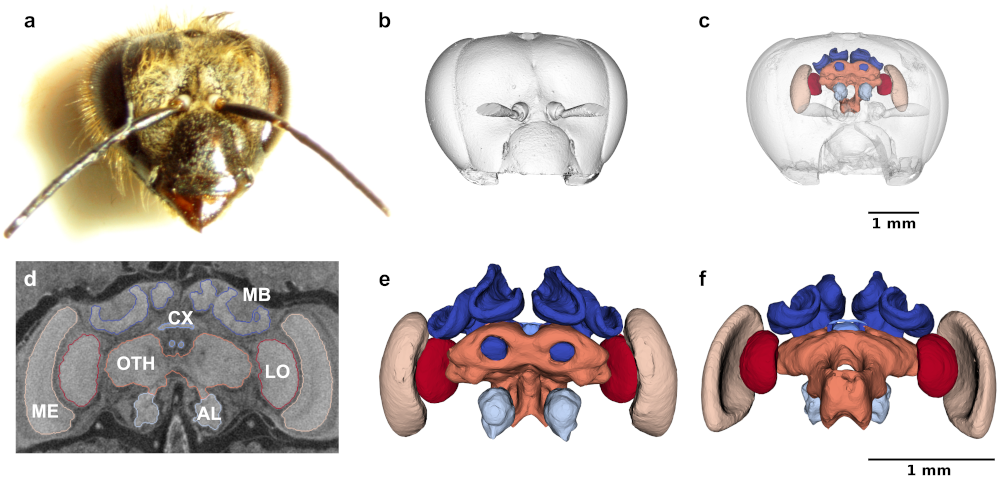
Large numbers of brain samples can reveal minor but statistically and biologically relevant variations that provide important insights into animal behavior, ecology, and evolution. Here, we used micro-CT imaging and Biomedisa's deep learning feature to perform automated analysis of 187 bee brains (honeybees and bumblebees). In bumblebees, we found a significantly larger right side of the optic and antennal lobes, providing a potential explanation for reported variations in visual and olfactory learning (bees learn better with their right eye and right antenna). Check out the paper!
Deep learning closes the gap to smart interpolation
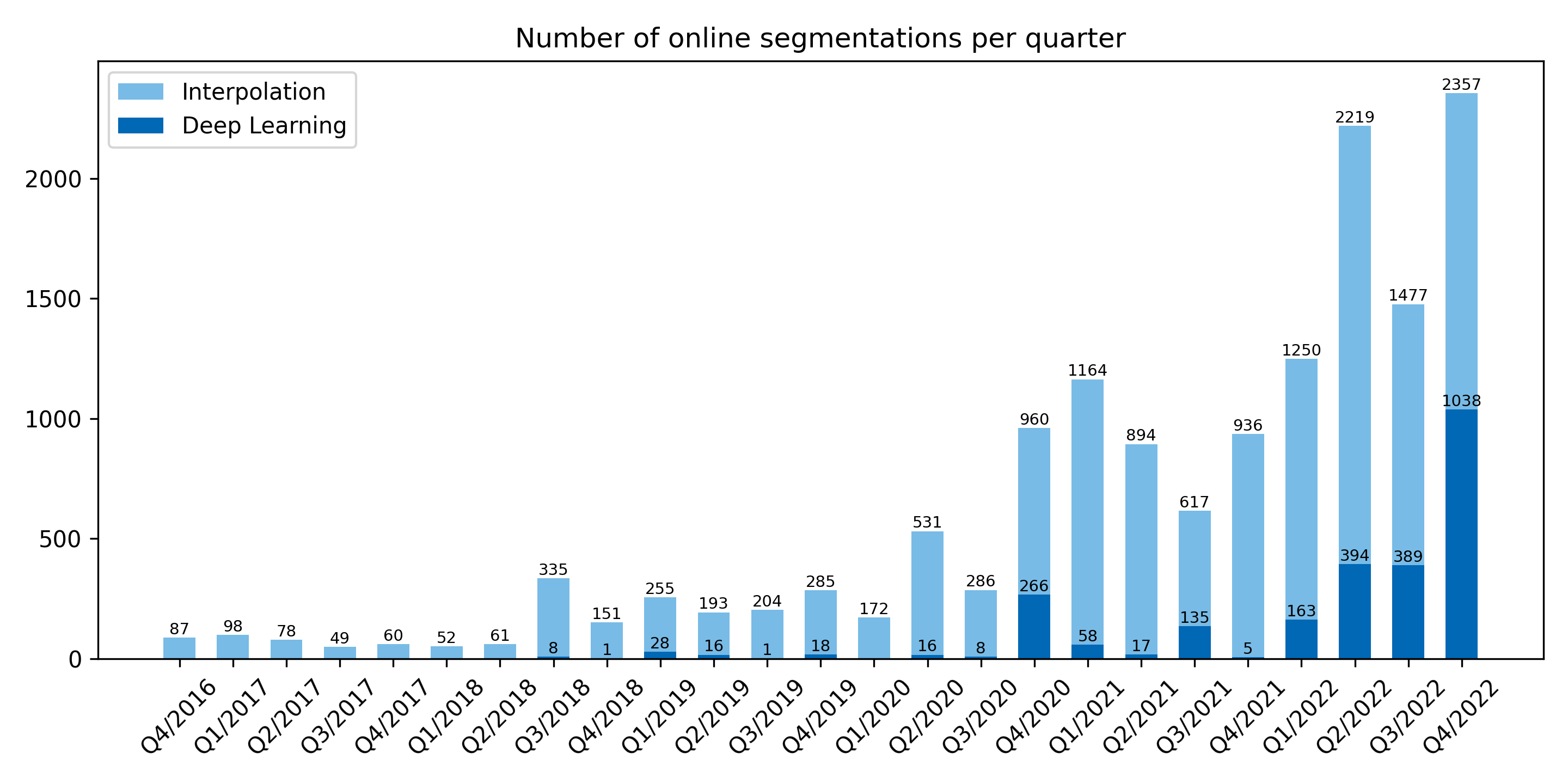
Posted on 03 jan 2023 by Philipp
Using Biomedisa's deep learning online is becoming more and more popular. The number of AI-based segmentations performed online exceeded three quarters of the interpolation tasks in the last three months of 2022. For those without access to GPUs, training your model online may be the optimal choice. Automated segmentation can then also achieved locally even with a CPU.
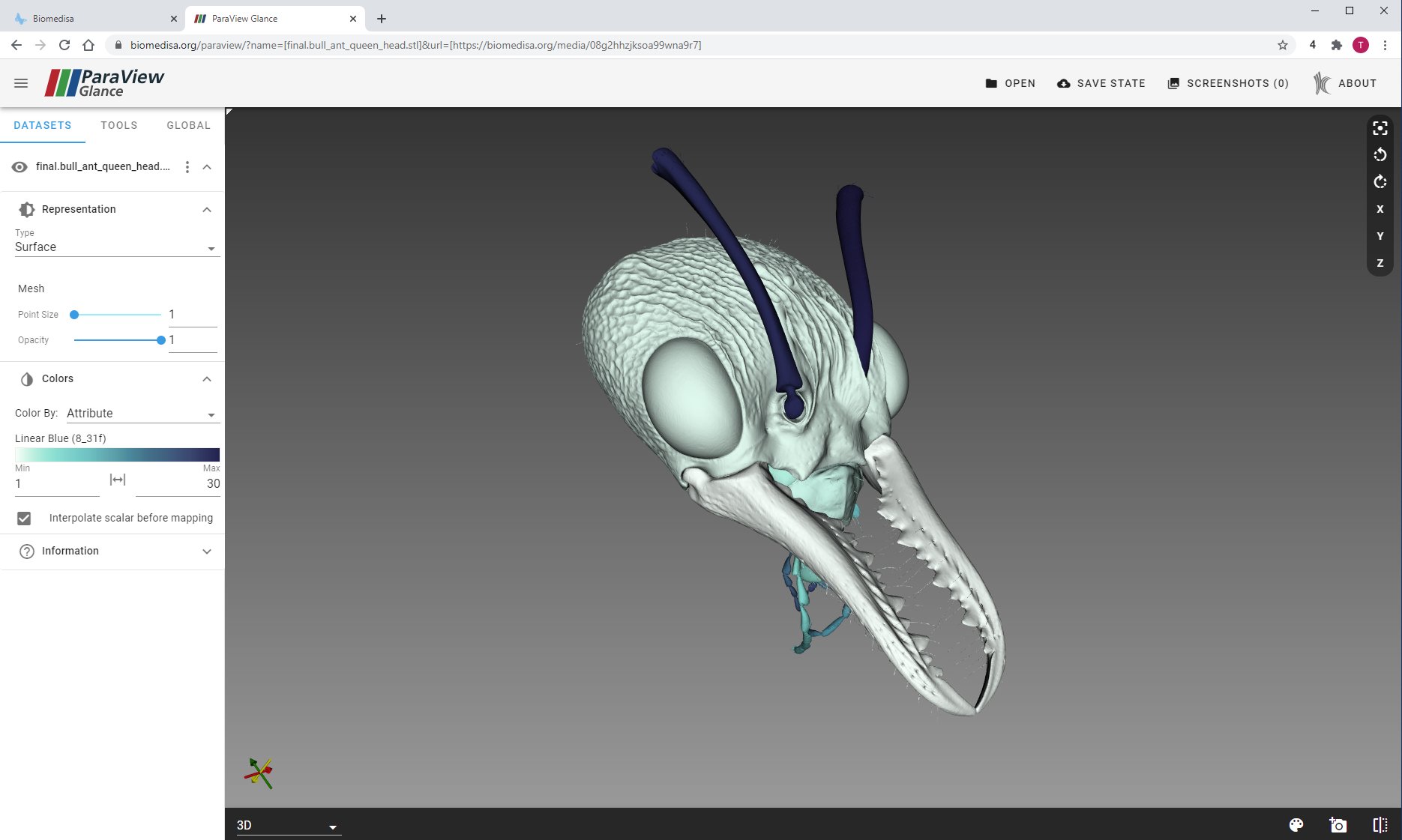
Biomedisa uses ParaView Glance for visualization
Posted on 22 nov 2020 by Philipp
Biomedisa enables volume rendering with ParaView Glance. Use the 3D feature button to render a selection of files. In addition, you can use the newly added mesh generator to visualize your segmentation results. Choose "Attribute" for "Color By" to color each segment individually. Check out the head of the bull ant queen.
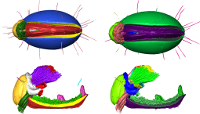
Tutorial: segmentation of a Trigonopterus weevil with Biomedisa
Posted on 09 nov 2020 by Philipp
This short tutorial shows how to use Biomedisa to segment a weevil from a tomographic image stack (presented by Thomas van de Kamp, KIT).
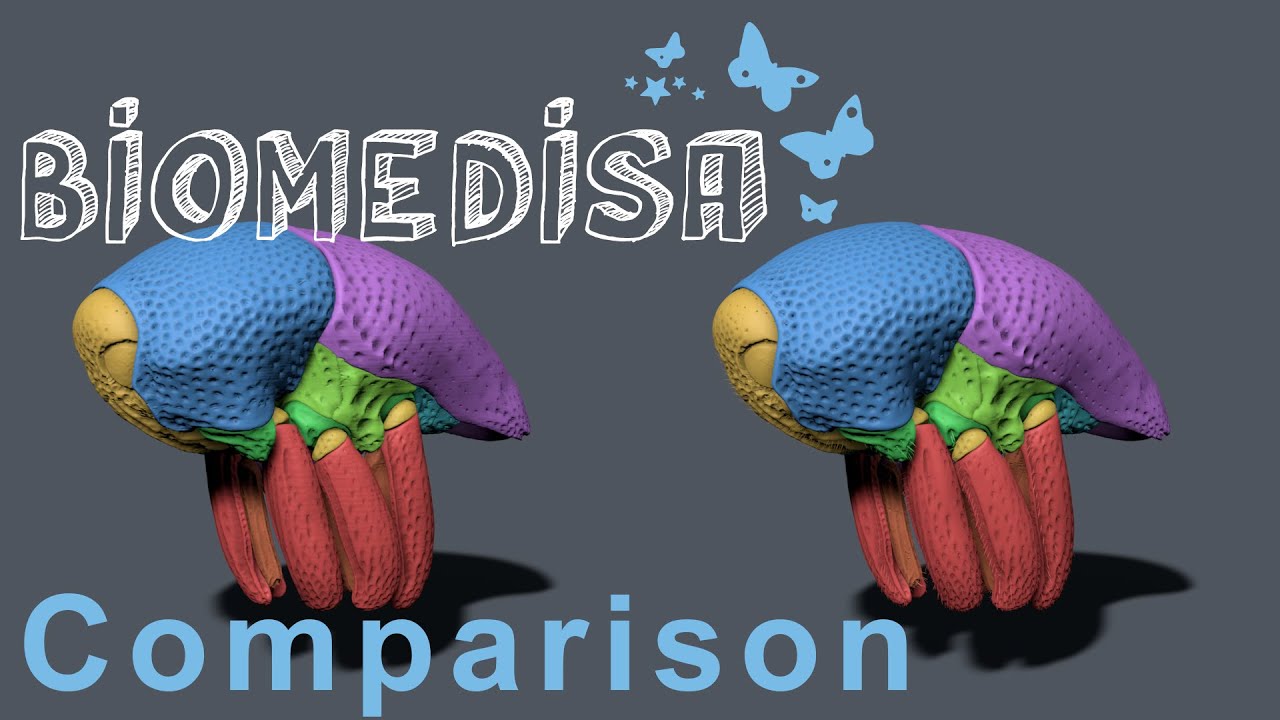
Comparison of Biomedisa with a conventional segmentation approach
Posted on 09 nov 2020 by Philipp
Comparison of Biomedisa with a conventional approach for segmenting a weevil from a tomographic image stack (Lösel et al. Nat. Commun.).
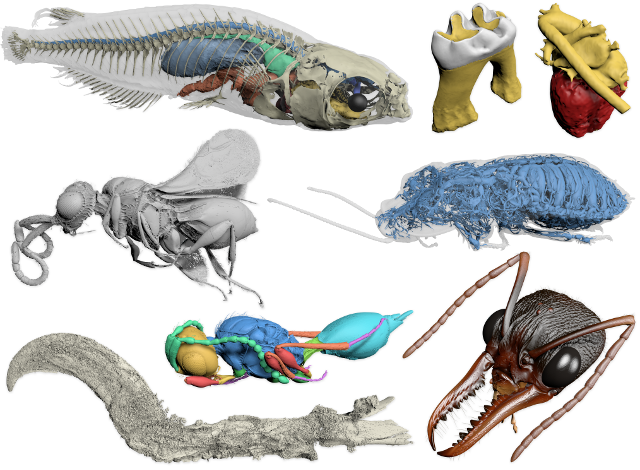
Biomedisa examples
Posted on 09 nov 2020 by Philipp
Biomedisa segmentation results of a medaka rice fish, a fossil parasoid wasp reconstructed from Baltic amber, the tracheal system of a hissing cockroach, and the head of a bull ant queen (featured in Lösel et al. Nat. Commun.).
Biomedisa has been published in Nature Communications
Posted on 04 nov 2020 by Philipp
Biomedisa has been published as an open-source project in Nature Communications. The paper demonstrates that Biomedisa can drastically reduce both the time and human effort required to segment large images when compared to the conventional approach of densely pre-segmented slices, as well as when compared to other segmentation tools. Follow us on Twitter and YouTube for updated news and content.
Segmentation of teeth and mandibular bone of the Ocelot (Leopardus pardalis)
Posted on 11 feb 2020 by Philipp
Data acquisition was performed at the School of Dentistry from a sample of the Biology Department (Universidad del Valle, Cali, Colombia). The segmentation was done with AVIZO and Biomedisa, the 3D rendering and animation with Dragonfly. (Video: Julián Balanta-Melo).
This is not a SEM micrograph!
Posted on 4 okt 2019 by Philipp
It’s a surface mesh based on a fast synchrotron microCT scan of an Ichneumonid wasp at IPS@KIT. Scan duration: 43s, pre-segmentation with Amira, semi-automated segmentation with Biomedisa, rendering with CINEMA 4D. (Video: Thomas van de Kamp).
From high-resolution μCT to the analysis of 3D reconstructions using Biomedisa
Posted on 12 mar 2019 by Philipp
Julián Balanta-Melo et al. studied the effect of masticatory function on tooth enamel and dentin in adult mouse molars after segmentation of three separate materials (enamel, dentin, and alveolar bone). Left: 3D reconstruction with enamel (white) and dentin (yellow) without surrounding alveolar bone tissue. Their results were published in Revista Estomatología.
Parasites discovered in fossil fly pupae
Posted on 28 aug 2018 by Philipp
Wasps in several-million-year-old fly pupae studied with synchrotron X-ray micro tomography and analysed with Biomedisa. Parasitic wasps existed as early as several million years ago. Within a project coordinated by Karlsruhe Institute of Technology (KIT), researchers of various disciplines for the first time definitively discovered fossil parasites inside their hosts. The scientists studied fly pupae from old collections using ultrafast X-ray imaging. They found 55 cases of parasitation and described four extinct wasp species that were unknown until now. Their findings are reported in Nature Communications. The data is available at fossils.kit.edu and in our gallery.
Digital resurgence of a parasitic wasp
Posted on 28 aug 2018 by Philipp
The parasitic wasp Xenomorphia resurrecta deposits an egg in a fly pupa. Following imaging of the mineralized fly pupae at the imaging beamline of the Institute for Photon Science and Synchrotron Radiation (IPS) at Karlsruhe Institute of Technology, the parasitic wasps from the Paleogene were reconstructed digitally with Biomedisa. (Video: Thomas van de Kamp, KIT; Nature Communications)
Biomedisa AI
Posted on 31 may 2018 by Philipp
Artificial intelligence in the form of deep neural networks has become an established method not only in speech recognition but also in image processing. Biomedisa AI is optimized for the segmentation of 3D image data. Artificial neural networks can be trained on fully pre-segmented data and applied to new, unknown data, enabling fully automated segmentation within seconds. See our gallery for an example.
DESY Photon Science Users' Meeting in Hamburg
Posted on 17 jan 2018 by Philipp
Users, collaborators and scientists interested in using photon sources for their research, will meet at the annual DESY Photon Science Users’ Meeting "Research with Synchrotron Radiation and FELs" from 25 June to 26 January in Hamburg. We'll present Biomedisa within the scope of the talk "The NOVA project: maximizing beam time efficiency through synergistic analyses of SRμCT data" in the satellite "Helmholtz-Zentrum Geesthacht GEMS Outstation: Materials Research and High Resolution Imaging". We are looking forward to seeing you in Hamburg.
Biomedisa supports Amira file format
Posted on 10 dec 2017 by Alejandra
To support the workflow of Amira users, it is no longer necessary to convert Amira files to TIFF format. Images and labels can be loaded directly into Biomedisa as Amira files. The Biomedisa result is then saved as an Amira mesh with all meta information and can be loaded back into Amira.
Biomedisa at ICTMS 2017 in Lund, Sweden
Posted on 18 may 2017 by Philipp
The ICTMS 2017, from 26 June to 30 June, will bring together an international group of scientists, from universities, research organisations and industry, to discuss a broad range of issues related to the use of 3D tomographic imaging in materials and structures. An abstract can be found here.
Biomedisa at ISC High Perfomance 2017 in Frankfurt
Posted on 17 jul 2017 by Philipp
The ISC High Performance 2017, from 19 June to 22 June, is dedicated to tackling HPC technological development and its application in scientific fields, as well as its adoption in commercial environments. It brings together researchers from academy and industry in the field of high performance computing. Biomedisa was represented together with the Computing Center of Heidelberg University and the EMCL.
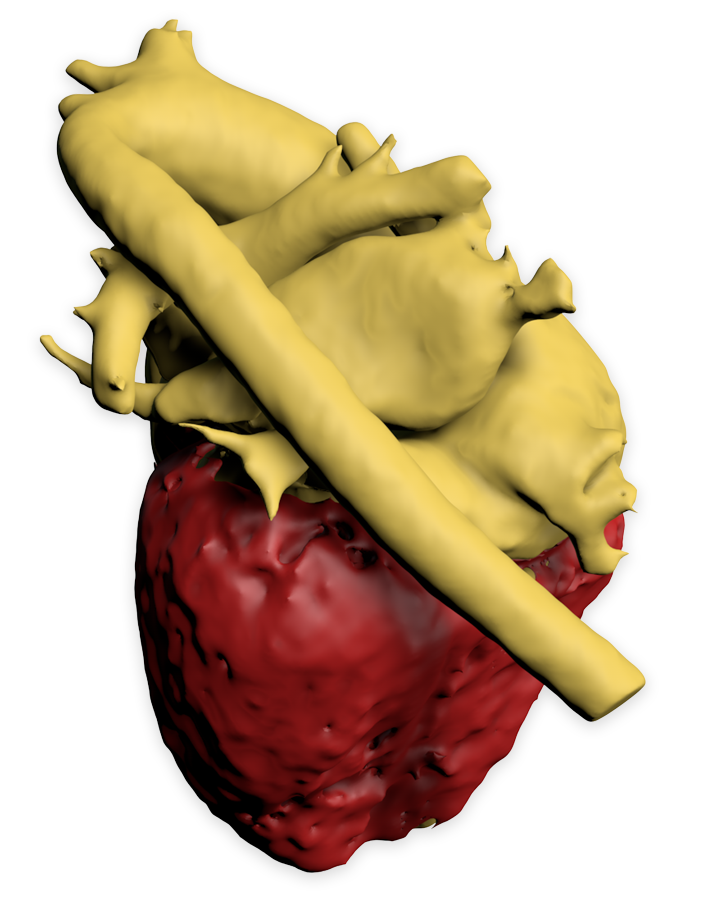
Biomedisa for Whole-Heart and Great Vessel Segmentation
Posted on 31 jan 2017 by Philipp
Segmenting the blood pool and myocardium from a 3D cardiovascular magnetic resonance image allows to create a patient-specific heart model for surgical planning in children with complex congenital heart disease. By using Biomedisa we achieved a high segmentation accuracy combined with a small amount of manual labeling and a short computing time.
3D reconstructions come to life
Posted on 30 jan 2017 by Philipp
Instead of using static images or movies, complex morphological 3D models based on segmented datasets have been published in the last years as 3D PDF files. 3D PDF files allow the user to handle and examine relevant structures interactively. As an example, you can download a 3D model of Euphthiracarus reticulatus by Sebastian Schmelzle here and open it with Adobe Reader. If you want to find out more, download more examples or even read about animated 3D PDF files, we strongly recommend you to read "Three-Dimensional Reconstructions Come to Life – Interactive 3D PDF Animations in Functional Morphology" by Thomas van de Kamp et al.
12. Modellierungstag Rhein-Neckar, 8. Dec 2016,
HGS MathComp, Heidelberg
Posted on 22 nov 2016 by Philipp
On 8 December 2016 the 12th Modellierungstag Rhein-Neckar will focus on data visualization. The event aims to enable an exchange between scientists, developers, theorists, and industry. We will present Biomedisa and its recent developments.
Biomedisa at MICCAI 2016 in Athens, Greece
Posted on 22 nov 2016 by Philipp
MICCAI 2016, the 19th International Conference on Medical Image Computing and Computer Assisted Intervention, was held from 17 October to 21 October in Athens, Greece. The annual MICCAI conference attracts world leading biomedical scientists, engineers, and clinicians from a wide range of disciplines associated with medical imaging and computer assisted intervention. The biomedical image segmentation app was presented in a workshop being held along with the conference.
Morphology Yesterday, Today and Tomorrow
Posted on 22 nov 2016 by Philipp
From 13 October to 16 October the 9th Graduiertentreffen der DZG (Deutsche Zoologische Gesellschaft e.V.) Fachgruppe Morphologie took place at KIT, Karlsruhe. Presentations were given about classical, modern, and future-oriented morphological image processing techniques. We presented our application and came into contact with many users and developers of morphological analysis tools. On 13 September we also gave a talk at the DZG Workshop "Engineering tools in morphology – automated image processing, rapid prototyping and determination of material properties" in Kiel, Germany.
Biomedisa at SPIE 2016 in San Diego, USA
Posted on 2 mar 2016 by Philipp
From 27 February to 3 March 2016 the SPIE Medical Imaging Conference 2016 took place in San Diego, California, USA. More than 1,000 presentations were given on the latest research in the area of medical imaging covering various topics such as Physics of Medical Imaging, Image Processing, Computer-Aided Diagnosis as well as Image-Guided Procedures, Robotic Interventions, and Modeling. Biomedisa presented its diffusion algorithm based on their research „Enhancing a Diffusion Algorithm for 4D Image Segmentation Using Local Information" within the BMBF Project ASTOR.